What is described in this manual is the full version, and so is likely to contain some perhaps more esoteric options that few people will need to use. For many operations it is convenient to be able to process sets of data together - for example to calculate a consensus sequence for a subset of the contigs. This introductory section also contains a complete list of the options in the gap4 main menus see section Gap4 Menus. One of the most powerful features of gap4 is its graphical user interface which enables the data to be viewed and manipulated at several levels of resolution. The displays which provide these different views are introduced, with several screenshots see section Introduction to the gap4 User Interface. For example it is easy to create a beginners version of gap4 which has only a subset of functions. The main topics that are introduced are listed in the current section. 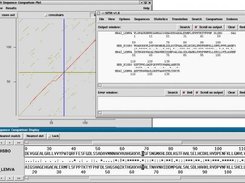
| Uploader: | Kazranris |
| Date Added: | 1 May 2015 |
| File Size: | 47.91 Mb |
| Operating Systems: | Windows NT/2000/XP/2003/2003/7/8/10 MacOS 10/X |
| Downloads: | 90165 |
| Price: | Free* [*Free Regsitration Required] |
The displays which provide these different views are introduced, with several screenshots see section Introduction to the gap4 User Interface.
Everybody employs a subset of options and has their favourite way of accessing and using them.
Staden Package
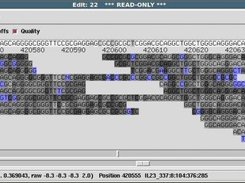
Generally the 3' ends of readings from sequencing stwden are of too low a quality to be used to create reliable consensus, but they can be useful, for example, for finding joins between contigs see section Use of the "hidden" poor quality data. The ability to annotate segments of readings and the consensus can be very convenient see section Annotating and masking readings and contigs.
Note that gap4 is a very flexible program, and is designed so that it can easily be configured to suit different purposes and ways of working. One of the most powerful features of gap4 is its graphical user interface which enables the data to be viewed and manipulated at several levels of resolution.
Staden Package Home
This introductory section also contains a ga4p list of the options in the gap4 main menus see section Gap4 Menus. In addition to sequence assembly, gap4 can be used for managing mutation study data and for helping to discover and check for mutations see section Introduction to Searching for Mutations. Although there is a lot of it, users are encouraged to go through the whole of the documentation once, just to discover what is possible, and the way that best suits their own work.
This chapter serves as a cross reference point, to give an overview of the program and to introduce some of the important ideas which it uses. It is important to understand the different files used by our sequence assembly software, and how the data is processed before it reaches gap4 see section Summary of the Files used and the Preprocessing Steps.
Two further useful facilities of gap4 are "Lists" and "Notes". A new DNA sequence assembly program. What is described in this manual is the full version, and so is likely to contain some perhaps more esoteric options that few people will need to use. Stacen many operations it is convenient to be able to process sets of data together - for example to calculate a consensus gwp4 for a subset of the contigs.
Gap4 - Gap4-Introduction
Gap4 is very big and powerful. The original version was described in Bonfield,J. We introduced the use of base call accuracy values for speeding up sequencing projects see section The use of numerical estimates of base calling accuracy.
The program contains all the tools that would be expected from an assembly program plus many unique features and a very easily used interface. For example it is easy to create a beginners version of gap4 which has only a subset of functions.
At the very least, the whole of this introductory chapter should be read, as in the long run, it will save time. The main topics that are introduced are listed in the current section.

No comments:
Post a Comment